Note
Go to the end to download the full example code.
Flow Cytometry Simulation: Full System Example#
This tutorial demonstrates a complete flow cytometry simulation using the FlowCyPy library. It models fluidics, optics, signal processing, and classification of multiple particle populations.
Steps Covered:#
Configure simulation parameters and noise models
Define laser source, flow cell geometry, and fluidics
Add synthetic particle populations
Set up detectors, amplifier, and digitizer
Simulate analog and digital signal acquisition
Apply triggering and peak detection
Classify particle events based on peak features
Step 1: Define Flow Cell and Fluidics#
from FlowCyPy.units import ureg
from FlowCyPy.fluidics import (
Fluidics,
FlowCell,
ScattererCollection,
populations,
SampleFlowRate,
SheathFlowRate,
)
from FlowCyPy.fluidics import distributions
flow_cell = FlowCell(
sample_volume_flow=SampleFlowRate.MEDIUM.value,
sheath_volume_flow=SheathFlowRate.MEDIUM.value,
width=400 * ureg.micrometer,
height=150 * ureg.micrometer,
)
scatterer_collection = ScattererCollection()
medium_refractive_index = distributions.Delta(1.33)
diameter_dist = distributions.RosinRammler(
scale=200 * ureg.nanometer,
shape=10,
)
ri_dist = distributions.Normal(
mean=1.44,
standard_deviation=0.002,
low_cutoff=1.33,
)
sampling_method = populations.ExplicitModel()
population_0 = populations.SpherePopulation(
name="Pop 0",
medium_refractive_index=medium_refractive_index,
concentration=1e10 * ureg.particle / ureg.milliliter,
diameter=diameter_dist,
refractive_index=ri_dist,
sampling_method=sampling_method,
)
diameter_dist = distributions.RosinRammler(
scale=30 * ureg.nanometer,
shape=50,
)
ri_dist = distributions.Normal(
mean=1.44,
standard_deviation=0.002,
low_cutoff=1.33,
)
population_1 = populations.SpherePopulation(
name="Pop 1",
medium_refractive_index=medium_refractive_index,
concentration=5e11 * ureg.particle / ureg.milliliter,
diameter=diameter_dist,
refractive_index=ri_dist,
sampling_method=populations.GammaModel(number_of_samples=5_000),
)
scatterer_collection.add_population(population_0, population_1)
scatterer_collection.dilute(factor=80)
fluidics = Fluidics(scatterer_collection=scatterer_collection, flow_cell=flow_cell)
Step 2: Define Optical Subsystem#
from FlowCyPy.opto_electronics import (
Detector,
Digitizer,
OptoElectronics,
Amplifier,
source,
circuits,
)
analog_processing = [
circuits.BaselineRestorationServo(time_constant=100 * ureg.microsecond),
circuits.BesselLowPass(cutoff_frequency=2 * ureg.megahertz, order=4, gain=2),
]
source = source.Gaussian(
waist_z=10e-6 * ureg.meter, # Beam waist along flow direction (z-axis)
waist_y=60e-6 * ureg.meter,
wavelength=405 * ureg.nanometer,
optical_power=200 * ureg.milliwatt,
rin=-140 * ureg.dB_per_Hz,
bandwidth=10 * ureg.megahertz,
)
detectors = [
Detector(
name="side",
phi_angle=90 * ureg.degree,
numerical_aperture=1.1,
responsivity=1 * ureg.ampere / ureg.watt,
),
Detector(
name="forward",
phi_angle=0 * ureg.degree,
numerical_aperture=0.3,
cache_numerical_aperture=0.1,
responsivity=1 * ureg.ampere / ureg.watt,
),
]
digitizer = Digitizer(
sampling_rate=60 * ureg.megahertz,
bit_depth=14,
use_auto_range=True,
channel_range_mode="shared",
)
amplifier = Amplifier(
gain=10 * ureg.volt / ureg.ampere,
bandwidth=10 * ureg.megahertz,
voltage_noise_density=0.0 * ureg.nanovolt / ureg.sqrt_hertz,
current_noise_density=0.0 * ureg.femtoampere / ureg.sqrt_hertz,
)
opto_electronics = OptoElectronics(
digitizer=digitizer,
detectors=detectors,
source=source,
amplifier=amplifier,
analog_processing=analog_processing,
)
Step 3: Signal Processing Configuration#
from FlowCyPy.digital_processing import (
DigitalProcessing,
peak_locator,
discriminator,
)
triggering = discriminator.FixedWindow(
trigger_channel="side",
threshold="4sigma",
pre_buffer=40,
post_buffer=40,
max_triggers=-1,
)
peak_algo = peak_locator.GlobalPeakLocator()
digital_processing = DigitalProcessing(
discriminator=triggering,
peak_algorithm=peak_algo,
)
Step 4: Run Simulation#
from FlowCyPy import FlowCytometer
cytometer = FlowCytometer(
fluidics=fluidics,
background_power=0.001 * ureg.milliwatt,
)
run_record = cytometer.run(
opto_electronics=opto_electronics,
digital_processing=digital_processing,
run_time=1 * ureg.millisecond,
)
_ = run_record.event_collection.plot(x="Diameter")
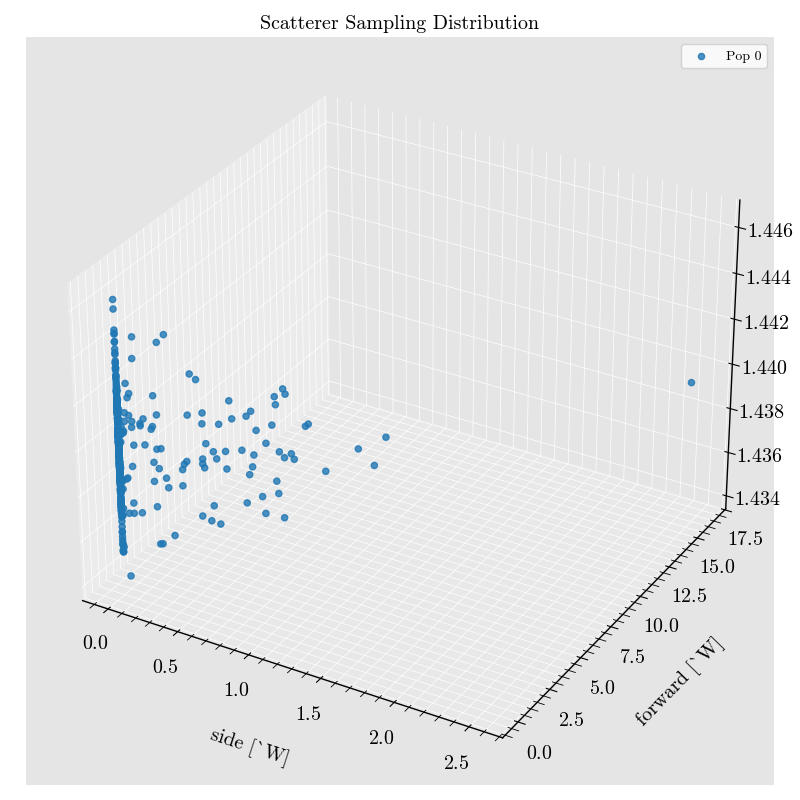
Step 5: Plot Events and Raw Analog Signals#
_ = run_record.event_collection.plot(x="forward")
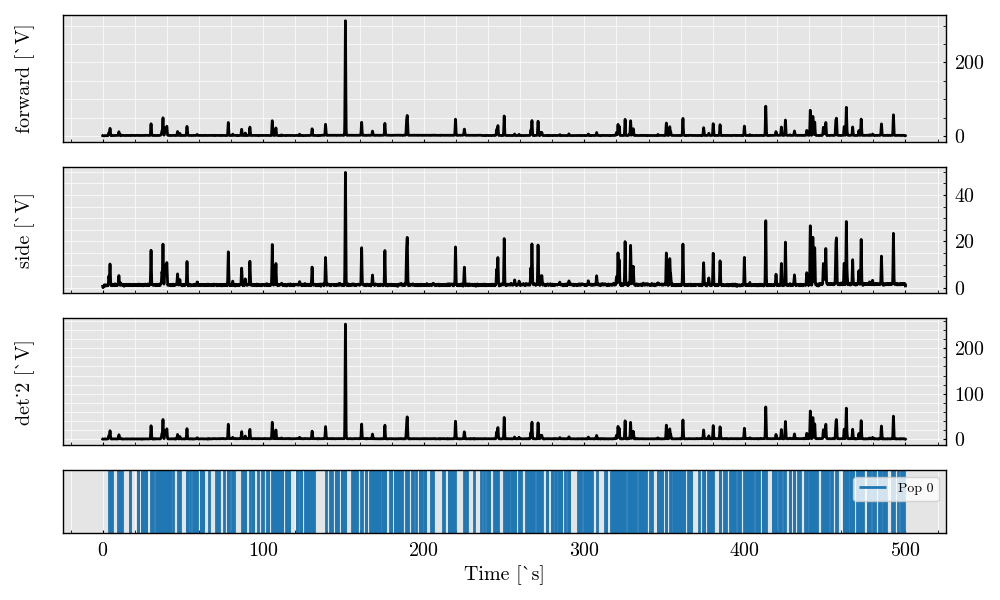
Plot raw analog signals#
_ = run_record.plot_analog(figure_size=(12, 8))
findfont: Font family ['cursive'] not found. Falling back to DejaVu Sans.
findfont: Generic family 'cursive' not found because none of the following families were found: Apple Chancery, Textile, Zapf Chancery, Sand, Script MT, Felipa, Comic Neue, Comic Sans MS, cursive
findfont: Font family ['Arial'] not found. Falling back to DejaVu Sans.
findfont: Font family ['Arial'] not found. Falling back to DejaVu Sans.
findfont: Font family ['Arial'] not found. Falling back to DejaVu Sans.
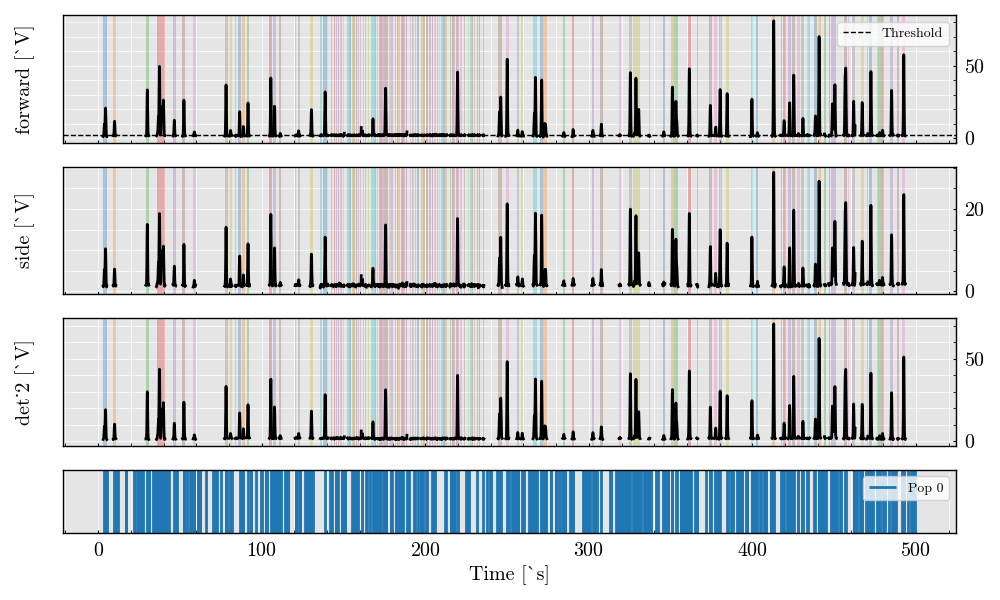
Step 6: Plot Triggered Analog Segments#
_ = run_record.plot_digital(figure_size=(12, 8))
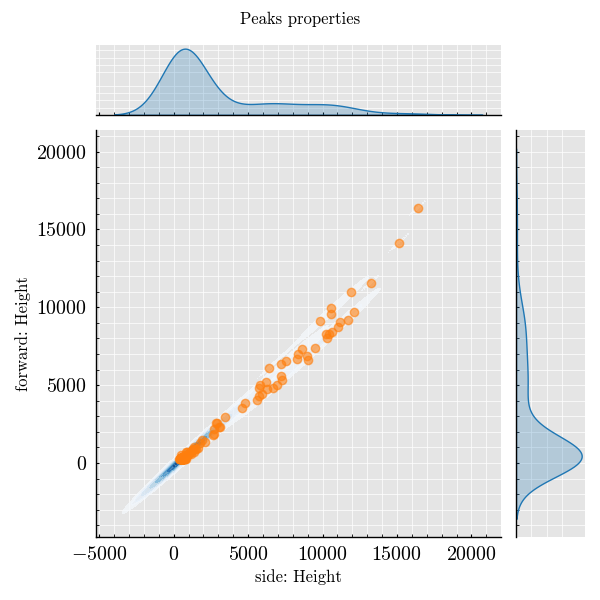
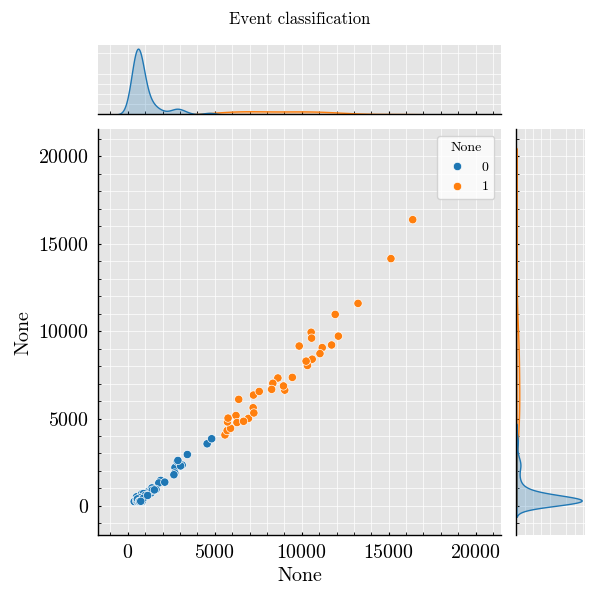
Step 7: Plot Peak Features#
_ = run_record.plot_peaks(x=("forward", "Height"))
Total running time of the script: (0 minutes 5.562 seconds)
