Note
Go to the end to download the full example code.
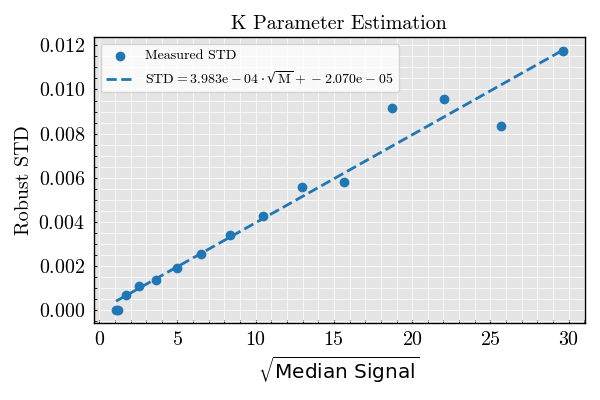
K Estimator Validation — Variable Bead Size, Fixed Illumination#
This example estimates the K parameter, which quantifies how the robust standard deviation scales with the square root of the signal strength under varying bead diameter.
The relationship is assumed to follow:
robust_std ≈ K * sqrt(median_signal)
We simulate a flow cytometry system with fixed illumination power and vary bead diameter across multiple runs.
Setup and configuration#
import matplotlib.pyplot as plt
import numpy as np
from FlowCyPy.units import ureg
from FlowCyPy import FlowCytometer
from FlowCyPy.fluidics import (
FlowCell,
Fluidics,
ScattererCollection,
SheathFlowRate,
SampleFlowRate,
)
from FlowCyPy.fluidics.populations import SpherePopulation
from FlowCyPy.opto_electronics import (
Digitizer,
circuits,
Detector,
OptoElectronics,
Amplifier,
source,
)
from FlowCyPy.digital_processing import (
DigitalProcessing,
peak_locator,
discriminator,
)
np.random.seed(3)
Helper functions#
def compute_signal_statistics(data: np.ndarray) -> dict:
"""
Compute robust and standard statistics from an event signal array.
"""
if data.size == 0:
raise ValueError("Signal array is empty.")
median = np.median(data)
percentile_84 = np.percentile(data, 84.13)
percentile_15 = np.percentile(data, 15.87)
mean = np.mean(data)
std = np.std(data)
robust_std = (percentile_84 - percentile_15) / 2.0
coefficient_of_variation = std / median if median != 0 else np.nan
robust_coefficient_of_variation = robust_std / median if median != 0 else np.nan
return {
"mean": mean,
"median": median,
"std": std,
"cv": coefficient_of_variation,
"robust_std": robust_std,
"robust_cv": robust_coefficient_of_variation,
}
def get_peaks(
flow_cytometer: FlowCytometer,
opto_electronics,
digital_processing,
run_time=5.0 * ureg.millisecond,
debug_mode=False,
):
"""
Run the flow cytometer and return the detected peaks.
"""
results = flow_cytometer.run(
opto_electronics=opto_electronics,
digital_processing=digital_processing,
run_time=run_time,
)
if debug_mode:
results.plot_analog()
plt.show()
results.plot_digital()
plt.show()
return results.peaks
def run_experiment(
flow_cytometer: FlowCytometer,
opto_electronics,
digital_processing,
bead_diameter,
illumination_power,
concentration,
run_time,
debug_mode=False,
):
"""
Configure the population and source power, then run one experiment.
"""
population = SpherePopulation(
name="population",
concentration=concentration,
diameter=bead_diameter,
medium_refractive_index=1.33,
refractive_index=1.47,
)
flow_cytometer.fluidics.scatterer_collection.populations = [population]
opto_electronics.source.optical_power = illumination_power
return get_peaks(
flow_cytometer=flow_cytometer,
opto_electronics=opto_electronics,
digital_processing=digital_processing,
run_time=run_time,
debug_mode=debug_mode,
)
def estimate_k(medians: np.ndarray, robust_stds: np.ndarray):
"""
Estimate K from a linear fit of robust_std versus sqrt(median).
"""
if medians.size < 2:
raise ValueError("At least two measurements are required to estimate K.")
x = np.sqrt(medians)
y = robust_stds
slope, intercept = np.polyfit(x, y, 1)
return slope, intercept
def plot_k_estimation(medians: np.ndarray, robust_stds: np.ndarray):
"""
Plot robust STD versus sqrt(median) and its linear fit.
"""
if medians.size < 2:
raise ValueError("At least two measurements are required to plot K estimation.")
x = np.sqrt(medians)
y = robust_stds
slope, intercept = np.polyfit(x, y, 1)
x_fit = np.linspace(np.min(x), np.max(x), 200)
y_fit = slope * x_fit + intercept
figure, axes = plt.subplots(figsize=(6, 4))
axes.scatter(x, y, label="Measured robust STD", zorder=3)
axes.plot(
x_fit,
y_fit,
"--",
label=rf"$STD = {slope:.3e} \cdot \sqrt{{M}} + {intercept:.3e}$",
zorder=2,
)
axes.set(
xlabel=r"$\sqrt{\mathrm{Median\ Signal}}$",
ylabel="Robust STD",
title="K Parameter Estimation",
)
axes.legend()
return figure, slope, intercept
Construct simulation components#
flow_cell = FlowCell(
sample_volume_flow=SampleFlowRate.MEDIUM.value,
sheath_volume_flow=SheathFlowRate.MEDIUM.value,
width=177 * ureg.micrometer,
height=433 * ureg.micrometer,
event_scheme="linear",
perfectly_aligned=True,
)
scatterer_collection = ScattererCollection()
fluidics = Fluidics(
scatterer_collection=scatterer_collection,
flow_cell=flow_cell,
)
illumination_source = source.Gaussian(
waist_z=10 * ureg.micrometer,
waist_y=60 * ureg.micrometer,
wavelength=450 * ureg.nanometer,
optical_power=200 * ureg.milliwatt,
include_shot_noise=True,
include_rin_noise=False,
)
digitizer = Digitizer(
bit_depth=24,
min_voltage=0 * ureg.volt,
max_voltage=100 * ureg.millivolt,
sampling_rate=60 * ureg.megahertz,
)
amplifier = Amplifier(
gain=10 * ureg.volt / ureg.ampere,
bandwidth=20 * ureg.megahertz,
voltage_noise_density=0 * ureg.volt / ureg.sqrt_hertz,
current_noise_density=0 * ureg.ampere / ureg.sqrt_hertz,
)
detector_0 = Detector(
name="default",
phi_angle=0 * ureg.degree,
numerical_aperture=0.5,
cache_numerical_aperture=0.0,
responsivity=1 * ureg.ampere / ureg.watt,
dark_current=0 * ureg.ampere,
)
baseline_restoration = circuits.BaselineRestorationServo(
time_constant=50 * ureg.microsecond
)
opto_electronics = OptoElectronics(
digitizer=digitizer,
analog_processing=[baseline_restoration],
detectors=[detector_0],
source=illumination_source,
amplifier=amplifier,
)
peak_algorithm = peak_locator.GlobalPeakLocator()
dynamic_window_discriminator = discriminator.DynamicWindow(
trigger_channel="default",
threshold=2 * ureg.microvolt,
pre_buffer=200,
post_buffer=200,
)
digital_processing = DigitalProcessing(
peak_algorithm=peak_algorithm,
discriminator=dynamic_window_discriminator,
)
flow_cytometer = FlowCytometer(
fluidics=fluidics,
background_power=illumination_source.optical_power * 0.00,
)
Run K estimation simulation#
debug_mode = False
run_time = 10 * ureg.millisecond
illumination_power = 200 * ureg.milliwatt
concentration = 2.5e7 * ureg.particle / ureg.milliliter
bead_diameters = np.linspace(300, 1500, 25) * ureg.nanometer
medians = []
robust_stds = []
robust_cvs = []
valid_bead_diameters = []
for index, bead_diameter in enumerate(bead_diameters):
print(
f"[INFO] Running simulation {index + 1}/{len(bead_diameters)} "
f"at bead diameter {bead_diameter}"
)
peaks = run_experiment(
flow_cytometer=flow_cytometer,
opto_electronics=opto_electronics,
digital_processing=digital_processing,
bead_diameter=bead_diameter,
illumination_power=illumination_power,
concentration=concentration,
run_time=run_time,
debug_mode=debug_mode,
)
if "Height" not in peaks:
raise KeyError("The peaks dataframe does not contain a 'Height' column.")
signal_array = peaks["Height"].values.astype(float)
if signal_array.size < 3:
print(
f"[WARNING] Skipping {bead_diameter} because only "
f"{signal_array.size} peak(s) were detected."
)
continue
statistics = compute_signal_statistics(signal_array)
medians.append(statistics["median"])
robust_stds.append(statistics["robust_std"])
robust_cvs.append(statistics["robust_cv"])
valid_bead_diameters.append(bead_diameter)
if debug_mode:
print(f"[DEBUG] Median: {statistics['median']:.3e}")
print(f"[DEBUG] Robust STD: {statistics['robust_std']:.3e}")
print(f"[DEBUG] Robust CV: {statistics['robust_cv']:.5f}")
medians = np.asarray(medians, dtype=float)
robust_stds = np.asarray(robust_stds, dtype=float)
robust_cvs = np.asarray(robust_cvs, dtype=float)
if medians.size < 2:
raise ValueError("Not enough valid measurements were collected to estimate K.")
[INFO] Running simulation 1/25 at bead diameter 300.0 nanometer
[INFO] Running simulation 2/25 at bead diameter 350.0 nanometer
[INFO] Running simulation 3/25 at bead diameter 400.0 nanometer
[INFO] Running simulation 4/25 at bead diameter 450.0 nanometer
[INFO] Running simulation 5/25 at bead diameter 500.0 nanometer
[INFO] Running simulation 6/25 at bead diameter 550.0 nanometer
[INFO] Running simulation 7/25 at bead diameter 600.0 nanometer
[INFO] Running simulation 8/25 at bead diameter 650.0 nanometer
[INFO] Running simulation 9/25 at bead diameter 700.0 nanometer
[INFO] Running simulation 10/25 at bead diameter 750.0 nanometer
[INFO] Running simulation 11/25 at bead diameter 800.0 nanometer
[INFO] Running simulation 12/25 at bead diameter 850.0 nanometer
[INFO] Running simulation 13/25 at bead diameter 900.0 nanometer
[INFO] Running simulation 14/25 at bead diameter 950.0 nanometer
[INFO] Running simulation 15/25 at bead diameter 1000.0 nanometer
[INFO] Running simulation 16/25 at bead diameter 1050.0 nanometer
[INFO] Running simulation 17/25 at bead diameter 1100.0 nanometer
[INFO] Running simulation 18/25 at bead diameter 1150.0 nanometer
[INFO] Running simulation 19/25 at bead diameter 1200.0 nanometer
[INFO] Running simulation 20/25 at bead diameter 1250.0 nanometer
[INFO] Running simulation 21/25 at bead diameter 1300.0 nanometer
[INFO] Running simulation 22/25 at bead diameter 1350.0 nanometer
[INFO] Running simulation 23/25 at bead diameter 1400.0 nanometer
[INFO] Running simulation 24/25 at bead diameter 1450.0 nanometer
[INFO] Running simulation 25/25 at bead diameter 1500.0 nanometer
Estimate K#
k_slope, k_intercept = estimate_k(
medians=medians,
robust_stds=robust_stds,
)
k_estimate = k_slope * ureg.AU**0.5
print("")
print("K estimation results")
print(f"Estimated K: {k_estimate}")
print(f"Linear intercept: {k_intercept:.6e}")
K estimation results
Estimated K: 0.23453034922330385 dimensionless
Linear intercept: -2.110897e+01
Plot estimation and diagnostics#
figure_k, _, _ = plot_k_estimation(
medians=medians,
robust_stds=robust_stds,
)
plt.show()
Total running time of the script: (1 minutes 36.719 seconds)
