Note
Go to the end to download the full example code.
Experiment Photodiode Angular Weights#
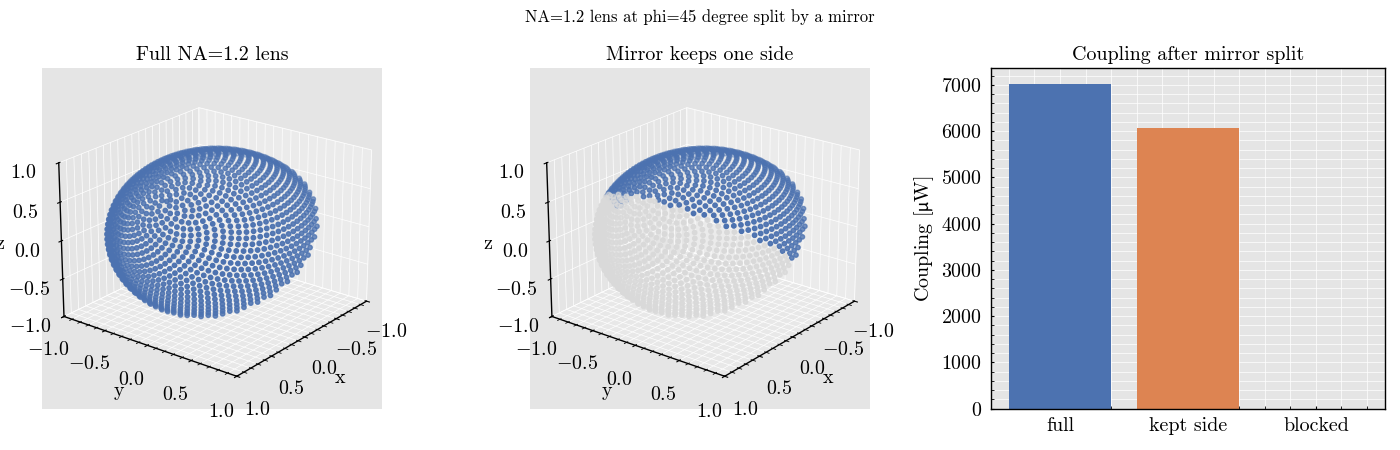
This example demonstrates how to model a large lens with NA=1.2 oriented
at phi=45 degree and split by a mirror plane. Only one side of the lens is
kept by building explicit angular_weights arrays for a sequential
PhotodiodeSet experiment.
At the moment, experiment-side angular weights are supported only through
PhotodiodeSet.build_sequential(...).
import numpy as np
import matplotlib.pyplot as plt
from PyMieSim.units import ureg
from PyMieSim.polarization import PolarizationState
from PyMieSim.single.source import Gaussian as SingleGaussian
from PyMieSim.single.scatterer import Sphere as SingleSphere
from PyMieSim.single.detector import Photodiode as SinglePhotodiode
from PyMieSim.experiment import Setup
from PyMieSim.experiment.detector_set import PhotodiodeSet
from PyMieSim.experiment.scatterer_set import SphereSet
from PyMieSim.experiment.source_set import GaussianSet
from PyMieSim.experiment.polarization_set import PolarizationSet
sampling = 1400
detector_numerical_aperture = 1.2
detector_phi_offset = 45 * ureg.degree
detector_gamma_offset = 0 * ureg.degree
detector_medium = 1.5
def normalize(vector: np.ndarray) -> np.ndarray:
return vector / np.linalg.norm(vector)
# Build one reference single detector so the mesh ordering can be used to create
# explicit experiment-side angular weight vectors.
single_source = SingleGaussian(
wavelength=1000 * ureg.nanometer,
polarization=PolarizationState(angle=0 * ureg.degree),
optical_power=1 * ureg.watt,
numerical_aperture=0.2,
)
single_scatterer = SingleSphere(
diameter=1000 * ureg.nanometer,
material=1.5 + 0.5j,
medium=1.0,
)
reference_detector = SinglePhotodiode(
numerical_aperture=detector_numerical_aperture,
phi_offset=detector_phi_offset,
gamma_offset=detector_gamma_offset,
sampling=sampling,
medium=detector_medium,
)
single_scatterer.init(single_source)
reference_detector.initialize_mesh(single_scatterer)
x_coordinates = np.asarray(reference_detector.mesh.cartesian.x.magnitude)
y_coordinates = np.asarray(reference_detector.mesh.cartesian.y.magnitude)
z_coordinates = np.asarray(reference_detector.mesh.cartesian.z.magnitude)
mesh_coordinates = np.column_stack([x_coordinates, y_coordinates, z_coordinates])
# Build a detector-local basis. The mirror plane contains the collection axis
# and the local horizontal direction, so the sign of the local vertical
# projection selects which side of the split lens is kept.
collection_axis = normalize(mesh_coordinates.mean(axis=0))
reference_up = np.array([0.0, 0.0, 1.0])
if np.abs(np.dot(collection_axis, reference_up)) > 0.95:
reference_up = np.array([0.0, 1.0, 0.0])
local_horizontal = normalize(np.cross(reference_up, collection_axis))
local_vertical = normalize(np.cross(collection_axis, local_horizontal))
local_vertical_coordinates = mesh_coordinates @ local_vertical
full_weights = np.ones(sampling, dtype=np.complex128)
mirror_kept_side = np.zeros(sampling, dtype=np.complex128)
mirror_kept_side[local_vertical_coordinates >= 0.0] = 1.0
blocked_weights = np.zeros(sampling, dtype=np.complex128)
source_set = GaussianSet.build_sequential(
target_size=3,
wavelength=[1000, 1000, 1000] * ureg.nanometer,
polarization=PolarizationSet(angles=[0, 0, 0] * ureg.degree),
optical_power=[1, 1, 1] * ureg.watt,
numerical_aperture=[0.2, 0.2, 0.2],
)
scatterer_set = SphereSet.build_sequential(
target_size=3,
diameter=[1000, 1000, 1000] * ureg.nanometer,
material=[1.5 + 0.5j, 1.5 + 0.5j, 1.5 + 0.5j],
medium=[1.0, 1.0, 1.0],
)
detector_set = PhotodiodeSet.build_sequential(
target_size=3,
numerical_aperture=[detector_numerical_aperture] * 3,
phi_offset=[45, 45, 45] * ureg.degree,
gamma_offset=[0, 0, 0] * ureg.degree,
sampling=[sampling, sampling, sampling],
polarization_filter=[0, 0, 0] * ureg.degree,
medium=[detector_medium, detector_medium, detector_medium],
angular_weights=[
full_weights,
mirror_kept_side,
blocked_weights,
],
)
experiment = Setup(
scatterer_set=scatterer_set,
source_set=source_set,
detector_set=detector_set,
)
coupling = experiment.get_sequential("coupling")
figure = plt.figure(figsize=(14, 4.5))
axis_full = figure.add_subplot(1, 3, 1, projection="3d")
axis_full.scatter(x_coordinates, y_coordinates, z_coordinates, s=10, c="#4c72b0")
axis_full.set(title="Full NA=1.2 lens", xlabel="x", ylabel="y", zlabel="z")
axis_half = figure.add_subplot(1, 3, 2, projection="3d")
active_half = np.abs(mirror_kept_side) > 0.0
axis_half.scatter(
x_coordinates[active_half],
y_coordinates[active_half],
z_coordinates[active_half],
s=10,
c="#4c72b0",
)
axis_half.scatter(
x_coordinates[~active_half],
y_coordinates[~active_half],
z_coordinates[~active_half],
s=10,
c="#d9d9d9",
)
axis_half.set(title="Mirror keeps one side", xlabel="x", ylabel="y", zlabel="z")
for axis in (axis_full, axis_half):
axis.set_xlim(-1.0, 1.0)
axis.set_ylim(-1.0, 1.0)
axis.set_zlim(-1.0, 1.0)
axis.view_init(elev=22, azim=38)
axis_bar = figure.add_subplot(1, 3, 3)
axis_bar.bar(
["full", "kept side", "blocked"],
coupling * 1e6,
color=["#4c72b0", "#dd8452", "#9a9a9a"],
)
axis_bar.set(
title="Coupling after mirror split",
ylabel="Coupling [$\\mu$W]",
)
figure.suptitle("NA=1.2 lens at phi=45 degree split by a mirror")
figure.tight_layout()
if plt.get_backend().lower() != "agg":
plt.show()
Total running time of the script: (0 minutes 0.442 seconds)
